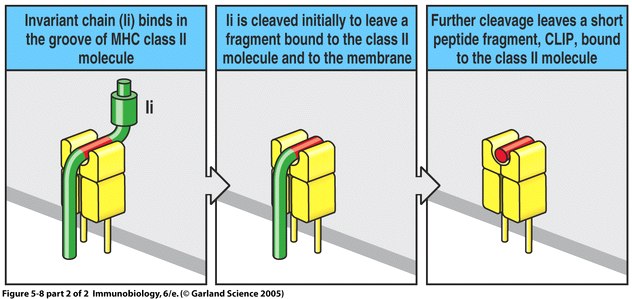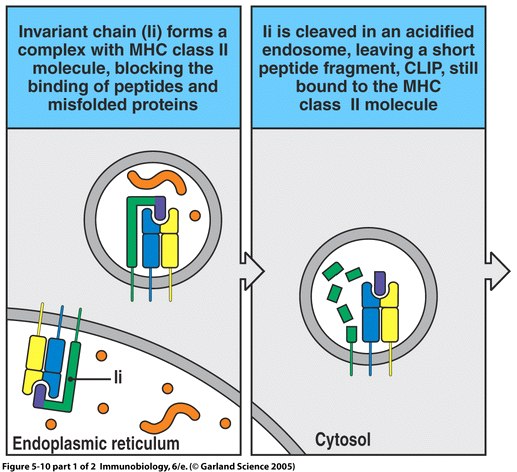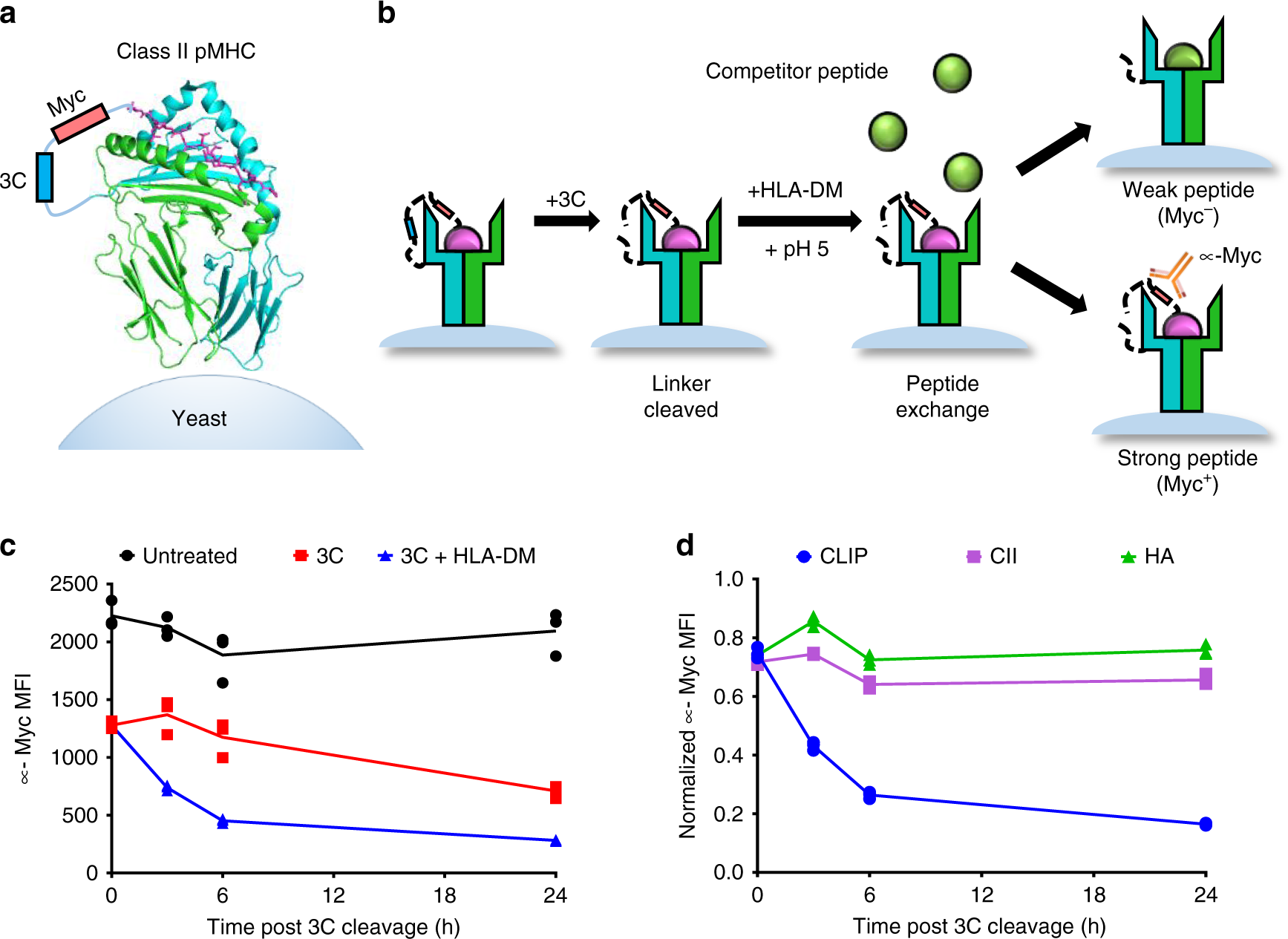
Repertoire-scale determination of class II MHC peptide binding via yeast display improves antigen prediction | Nature Communications

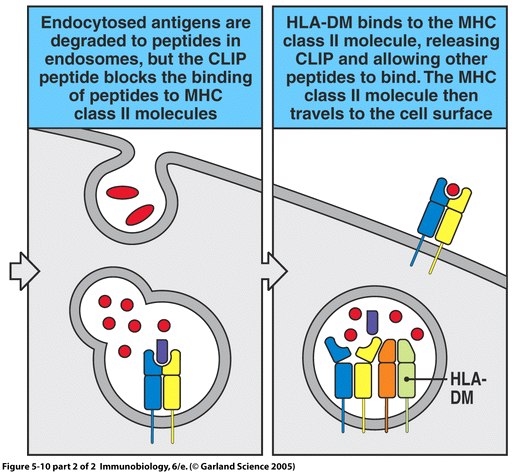
High-level intrinsic disorder explains the universality of CLIP binding to diverse MHC class II variants | Cellular & Molecular Immunology

High-resolution profiling of MHC II peptide presentation capacity reveals SARS-CoV-2 CD4 T cell targets and mechanisms of immune escape | Science Advances

Generation of MHC class II-peptide ligands for CD4 T-cell allorecognition of MHC class II molecules. - Abstract - Europe PMC

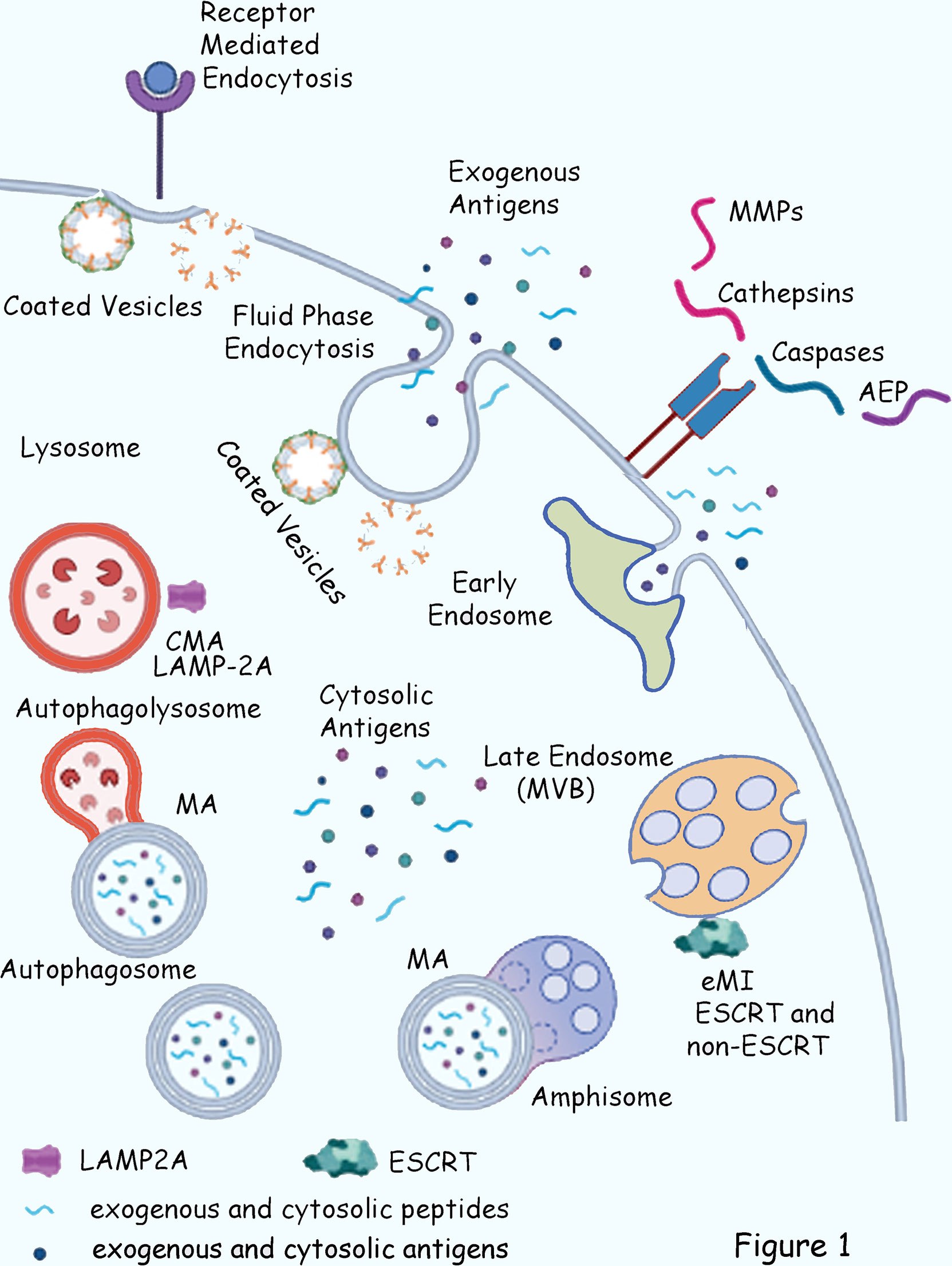
Intracellular localization of exogenous peptides and protein within the... | Download Scientific Diagram

Dynamic behavior of the HLA-DR1/CLIP 106-120 complex. (A) Close-up view... | Download Scientific Diagram

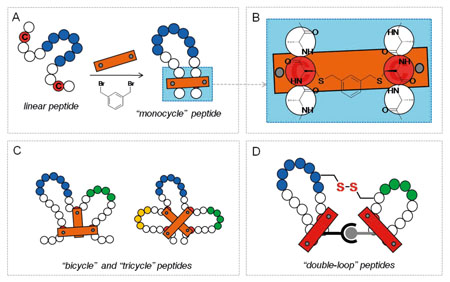
Deciphering the Structural Enigma of HLA Class-II Binding Peptides for Enhanced Immunoinformatics-based Prediction of Vaccine Epitopes | Journal of Proteome Research
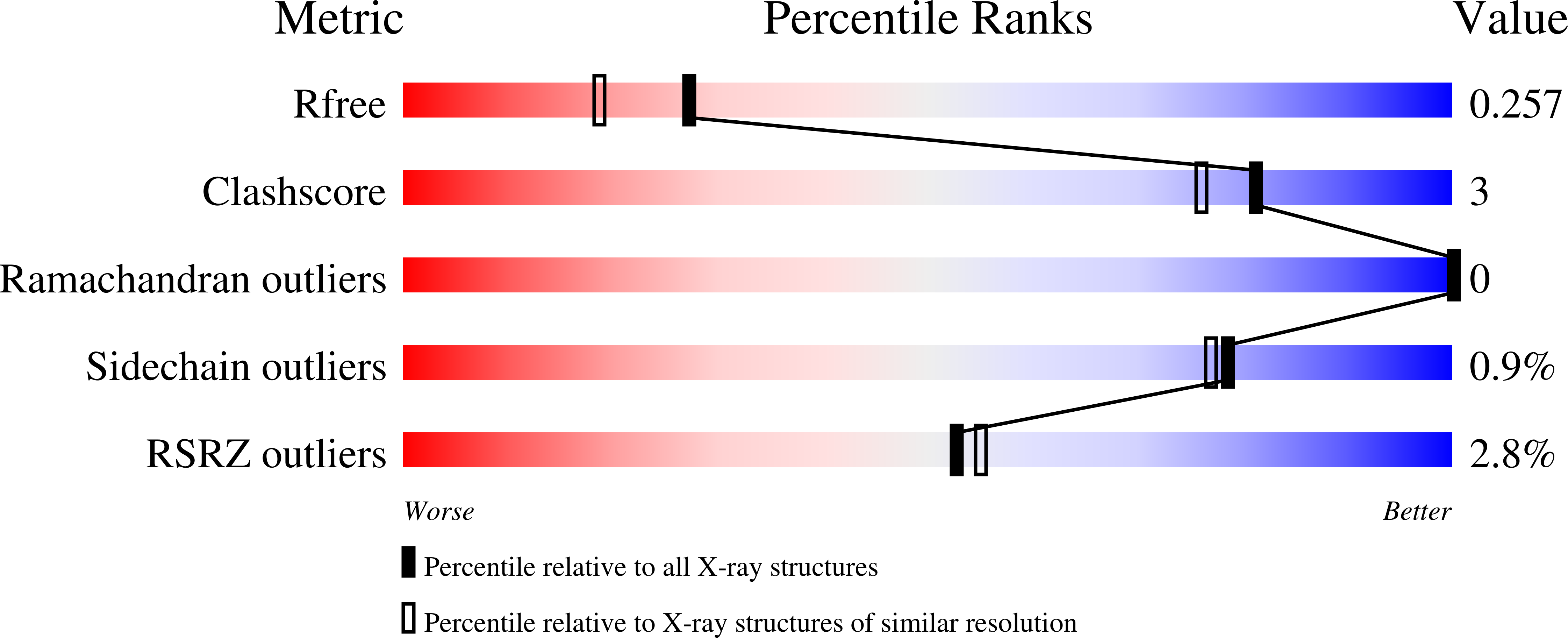
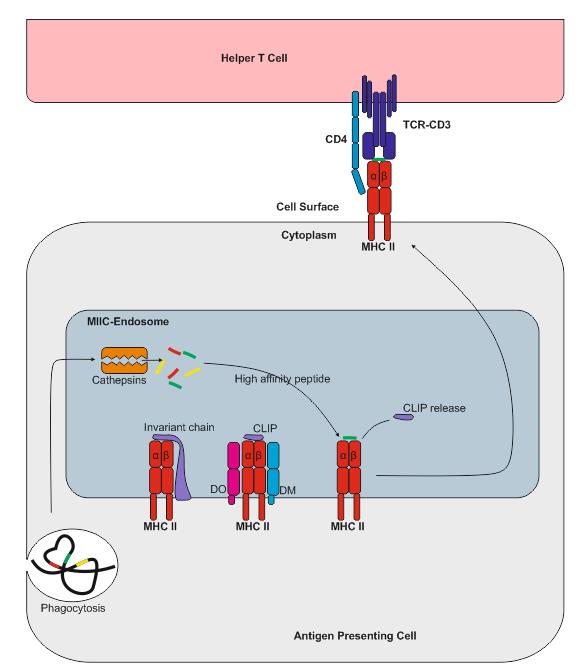
![PDF] Invariant chain, a chain of command. | Semantic Scholar PDF] Invariant chain, a chain of command. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/090358196965e735200921c37f7d927fe938721a/3-Figure3-1.png)